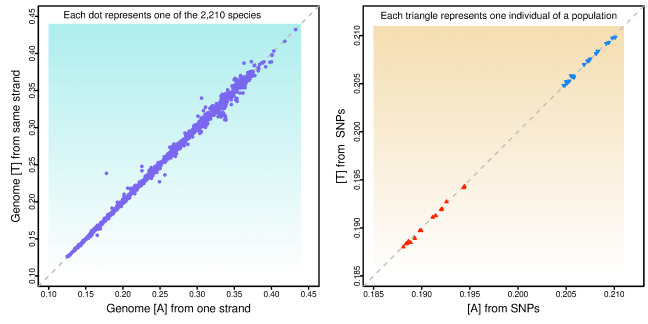
This graph demonstrates some of the genetic patterns uncovered in recent ISU research. The left panel shows the approximate equality of adenine and thymine in genomes across taxonomies. The right panel illustrates the same rule on the parts of genomes within species that vary among individuals and lead to divergence within a population. Courtesy of Jianming Yu. Larger image.
AMES, Iowa – A pair of genetics researchers at Iowa State University found striking patterns in the building blocks of DNA in a wide variety of species, according to their recently published paper.
The scientists said their work potentially offers new insight into how genes mutate over time and why some populations are more susceptible than others to certain diseases such as cancer.
Jianming Yu, an associate professor of agronomy and the Pioneer Distinguished Chair in Maize Breeding, and Xianran Li, a research assistant professor of agronomy, waded deep into the reams of data compiled on the genomes of 2,210 various species, including vertebrates, plants, fungi and bacteria. The research turned up some pronounced patterns, and it starts with counting the bases of chemical compounds that form strands of DNA.
“In a way, you could say this is about the genetics of genetics,” Yu said. “We always like to speculate on common mechanisms when we observe consistent patterns.”
DNA is made of chemical compounds known as nucleotide bases that pair together. Adenine matches with thymine, and guanine pairs with cytosine. Rather than looking at double helix structures, Li and Yu focused on single strands of DNA and ran statistical analyses on the preponderance of the four bases. They found that adenine-thymine and guanine-cytosine take up roughly equal proportions of genomes across all the taxonomies involved in the study.
Yu and Li ran similar analyses on multiple genomes within eight individual species for which such data exists, including humans. In those cases, they focused on parts of the genomes that vary among individuals. The researchers found what Yu called a “shocking consistency” regarding the approximate equality of the proportions of adenine-thymine and guanine-cytosine.
The research team found that individuals in a species with “bottlenecks” – events during which a small part of a population becomes separated and evolves independently – tend to have a higher proportion of adenine-thymine content in their genomes than individuals that descended from the population that didn’t go through the bottleneck.
The genomes of humans from Africa, where modern humans originate, show a higher proportion of guanine-cytosine content than human populations that migrated away from Africa over the millennia, Yu said. The researchers found similar results in other animal species and in plants.
These patterns may help us understand why certain populations experience higher rates of certain diseases, he said. For instance, the patterns may hint at which populations are likely to have genes that influence the development of breast cancer or the genes that govern sensitivity to ultraviolet light.
While the voluminous amounts of genome data used in the study already existed, Yu said his team was the first to analyze the data in this way.
“We found a fresh angle by integrating and mining data from many sources,” he said.
The research was published in Nucleic Acids Research, a peer-reviewed academic journal. Michael Scanlon, a professor of genetics at Cornell University, was a coauthor of the paper.