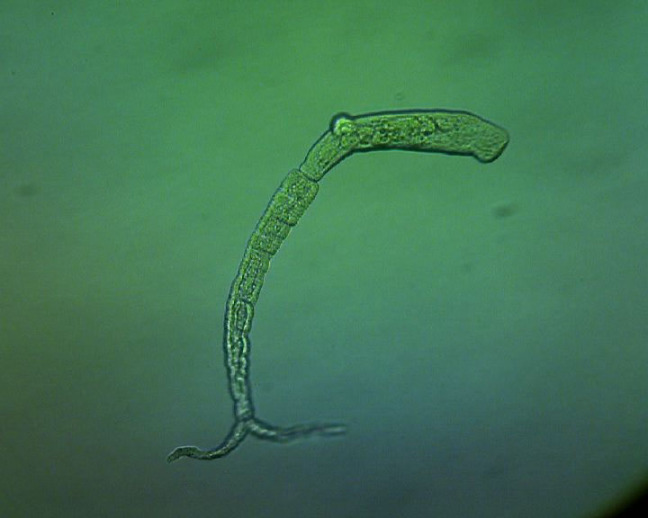
A schistosome cercaria, which is the form that invades human skin to infect more than 200 million people worldwide. Image courtesy of Sreemoyee Acharya. Larger image.
AMES, Iowa – Parasitic worms pose a major global health threat to both humans and animals, but a major study that sequenced the genomes of scores of worms could lead to new and more efficient ways to treat the illnesses they cause.
Tim Day, professor biomedical sciences in Iowa State University’s College of Veterinary Medicine, co-authored the study, published recently in the academic journal Nature Genetics. An international team of scientists conducted genomic studies of 81 worm species, including 45 that had never been sequenced before, and documented nearly a million new genes. With further study, some of those genes might present promising targets for new medical treatments to combat parasitic worms, Day said.
One in four humans carries a parasitic worm, he added, and infection rates are significantly higher in the developing world. River blindness and hookworm disease are perennial threats in tropical regions, and Day has spent years studying schistosomes, small flatworms that sicken more than 200 million people every year, according to the World Health Organization. In the animal world, the agricultural and veterinary industries continue to look for better ways to protect pets and livestock.
The comparative genomics study, the largest to date, looks at both nematodes, or roundworms, and platyhelminthes, or flatworms. The research identified gene families associated with critical functions for the worms, including genes that allow the parasites to overcome the immune responses of hosts, enable the worms to travel through host tissues and allow the parasites to feed. New treatments that impair those gene families could combat the diseases they cause, Day said.
The researchers found that some of those gene families show up in more than one species, meaning a single treatment could be effective against multiple parasites, spanning roundworms and flatworms.
“Not very long ago, the idea of having the entire genome of a single worm available to us was pretty awesome,” Day said. “Now, in one publication, we have the data from 81 different worms. The sheer volume and amplitude of this work gives us an opportunity to find commonalities among different species that was never possible before.”
Nic Wheeler, a former ISU graduate student in Day’s lab and a current postdoctoral researcher at the University of Wisconsin–Madison, searched the newly sequenced genomes for the presence and abundance of proteins that have served as successful therapeutic targets for a wide range of other infectious diseases. The new data reveal important genetic lines that are highly conserved among all parasitic worms, and some that are unique to just roundworms or flatworms.
“For decades, we and others have studied these historical drug targets without really understanding their breadth and conservation within the entirety of the flatworm and roundworm parasites,” Wheeler said. “This dataset gives us a much broader view of the biological importance of these targets, and it enables new avenues of inquiry into how we can exploit them for new therapies.”
Contacts
Timothy Day, Biomedical Sciences, 515-294-7100, day@iastate.edu
Fred Love, News Service, 515-294-0704, fredlove@iastate.edu
Quick look
An international team of scientists, including an ISU biomedical researcher, conducted genomic studies of 81 worm species, including 45 that had never been sequenced before, and documented nearly a million new genes. The research might identify promising targets for new medical treatments to combat parasitic worms, a major global threat to human and animal health.
Quote
“Not very long ago, the idea of having the entire genome of a single worm available to us was pretty awesome. Now, in one publication, we have the data from 81 different worms. The sheer volume and amplitude of this work gives us an opportunity to find commonalities among different species that was never possible before.”
Tim Day, professor of biomedical sciences